chloroplast
The chloroplast is the site of photosynthesis in eukaryotic cells, and is the site of the Calvin cycle just as the mitochondrion is the site of oxidative phosphorylation.
The thylakoid membrane, with its embedded photosystems, is the structural unit of photosynthesis. Both photosynthetic prokaryotes and eukaryotes possess membranes with embedded photosynthetic pigments. Only eukaryotes, which have a nuclear membrane and membrane-bound organelles, possess chloroplasts with an encapsulating membrane. The chloroplast has three compartments, while the mitochondrion has only two. Compartments within a chloroplast are the intermembranous space [3], the stroma [6], and the thylakoid lumen (8) within stromal and granal thylokoids [4,5].

1. outer membrane
2. inner membrane
3. intermembranous space
4. stromal thylakoid
5. granal thylakoid
6. stroma (cytosol)
7. granum (a stack of thylakoids)
8. internal lumen of granal and stromal thylakoids
(click to enlarge image)
The typical higher plant chloroplast is lenticular and approximately 5 microns at its largest dimension. Plant cells contain from 1 to 100 chloroplasts, depending on the type of cell. The mature chloroplast is typically bounded by outer (1) and inner (2) membranes that possess significantly different chemical constituents. In addition to enzymes that function in photosynthesis, chloroplasts also contain a circular DNA molecule and the protein-synthetic machinery characteristic of prokaryotes (below).
The smooth outer membrane (1) is freely permeable to molecules, and resembles the chemical constitution of the eukaryotic plasma membrane. The smooth inner membrane (2) contains many integral transporter proteins that regulate the passage of small molecules like sugars, and proteins (synthesized in the cytoplasm of the cell, but utilized within the chloroplast). The inner membrane chemically resembles prokaryotic cell membranes.
The folded thylakoid membranes perform the light reactions of photosynthesis utilizing Photosystems I and II, both of which include chlorophyll and carotenoid molecules. Thylakoid membranes also contain Cytochromes b and f and ATP synthase. The fluid stroma surrounding the thylakoid membranes includes the RUBISCO enzymes that perform the light-independent reactions of conversion of CO2 into organic molecules such as glucose. The stroma also includes the chloroplast genome (cpDNA genes) comprising many identical, circular molecules of DNA.
Each chloroplast contains about 40 to 80 grana (7), and each grana comprises about 5 to 30 thylakoids. The thylakoids are membranous disks about .25 to .8 microns in diameter, which contain protein complexes, pigments, and other accessory components. The phospholipid bilayer of the thylakoid is folded repeatedly into stacks of grana. (details) These stacks are connect by channels to form a single functional compartment – the internal lumen.
The thylakoid is the site of oxygenic photosynthesis in eukaryotic plants and algae, and in prokaryotic Cyanobacteria. Cyanobacteria possess thylakoid membranes, but as prokaryotes they do not contain chloroplasts. Chlorophyll, accessory pigments, and other integral membrane proteins transduce light energy to provide excited electrons (excitons) to electron transport chains, powering the formation of NADPH and ATP during photophosphorylation.
The current consensus is that chloroplasts originated from Cyanobacteria that have become endosymbionts. This is an origin analogous to the endosymbiotic origin of mitochondria, which are believed derived from "purple bacteria", alpha-proteobacteria most closely related to Rickettsiales.
The cpDNA genes encode some of the molecules needed for chloroplast function. Hundreds of others are transcribed from genes in cellular nucleus, translated into proteins in the cytoplasm, and transported into the chloroplast. It has been experimentally demonstrated that endosymbiotic engulfment of prokaryotes is followed by Endosymbiotic Gene Transfer.
Chloroplast RNA-binding and pentatricopeptide repeat proteins
Chloroplast gene expression is mainly regulated at the post-transcriptional level by numerous nuclear-encoded RNA-binding protein factors. In the present study, we focus on two RNA-binding proteins: cpRNP (chloroplast ribonucleoprotein) and PPR (pentatricopeptide repeat) protein. These are suggested to be major contributors to chloroplast RNA metabolism. Tobacco cpRNPs are composed of five different proteins containing two RNA-recognition motifs and an acidic N-terminal domain. The cpRNPs are abundant proteins and form heterogeneous complexes with most ribosome-free mRNAs and the precursors of tRNAs in the stroma. The complexes could function as platforms for various RNA-processing events in chloroplasts. It has been demonstrated that cpRNPs contribute to RNA stabilization, 3´-end formation and editing. The PPR proteins occur as a superfamily only in the higher plant species. They are predicted to be involved in RNA/DNA metabolism in chloroplasts or mitochondria. Nuclear-encoded HCF152 is a chloroplast-localized protein that usually has 12 PPR motifs. The null mutant of Arabidopsis, hcf152, is impaired in the 5´-end processing and splicing of petB transcripts. HCF152 binds the petB exon–intron junctions with high affinity. The number of PPR motifs controls its affinity and specificity for RNA. It has been suggested that each of the highly variable PPR proteins is a gene-specific regulator of plant organellar RNA metabolism.
T. Nakamura, G. Schuster, M. Sugiura and M. Sugita Chloroplast RNA-binding and pentatricopeptide repeat proteins Biochem. Soc. Trans.. (2004) 32, (571–574)
The thylakoid membrane, with its embedded photosystems, is the structural unit of photosynthesis. Both photosynthetic prokaryotes and eukaryotes possess membranes with embedded photosynthetic pigments. Only eukaryotes, which have a nuclear membrane and membrane-bound organelles, possess chloroplasts with an encapsulating membrane. The chloroplast has three compartments, while the mitochondrion has only two. Compartments within a chloroplast are the intermembranous space [3], the stroma [6], and the thylakoid lumen (8) within stromal and granal thylokoids [4,5].

1. outer membrane
2. inner membrane
3. intermembranous space
4. stromal thylakoid
5. granal thylakoid
6. stroma (cytosol)
7. granum (a stack of thylakoids)
8. internal lumen of granal and stromal thylakoids
(click to enlarge image)
The typical higher plant chloroplast is lenticular and approximately 5 microns at its largest dimension. Plant cells contain from 1 to 100 chloroplasts, depending on the type of cell. The mature chloroplast is typically bounded by outer (1) and inner (2) membranes that possess significantly different chemical constituents. In addition to enzymes that function in photosynthesis, chloroplasts also contain a circular DNA molecule and the protein-synthetic machinery characteristic of prokaryotes (below).
The smooth outer membrane (1) is freely permeable to molecules, and resembles the chemical constitution of the eukaryotic plasma membrane. The smooth inner membrane (2) contains many integral transporter proteins that regulate the passage of small molecules like sugars, and proteins (synthesized in the cytoplasm of the cell, but utilized within the chloroplast). The inner membrane chemically resembles prokaryotic cell membranes.
The folded thylakoid membranes perform the light reactions of photosynthesis utilizing Photosystems I and II, both of which include chlorophyll and carotenoid molecules. Thylakoid membranes also contain Cytochromes b and f and ATP synthase. The fluid stroma surrounding the thylakoid membranes includes the RUBISCO enzymes that perform the light-independent reactions of conversion of CO2 into organic molecules such as glucose. The stroma also includes the chloroplast genome (cpDNA genes) comprising many identical, circular molecules of DNA.
Each chloroplast contains about 40 to 80 grana (7), and each grana comprises about 5 to 30 thylakoids. The thylakoids are membranous disks about .25 to .8 microns in diameter, which contain protein complexes, pigments, and other accessory components. The phospholipid bilayer of the thylakoid is folded repeatedly into stacks of grana. (details) These stacks are connect by channels to form a single functional compartment – the internal lumen.
The thylakoid is the site of oxygenic photosynthesis in eukaryotic plants and algae, and in prokaryotic Cyanobacteria. Cyanobacteria possess thylakoid membranes, but as prokaryotes they do not contain chloroplasts. Chlorophyll, accessory pigments, and other integral membrane proteins transduce light energy to provide excited electrons (excitons) to electron transport chains, powering the formation of NADPH and ATP during photophosphorylation.
The current consensus is that chloroplasts originated from Cyanobacteria that have become endosymbionts. This is an origin analogous to the endosymbiotic origin of mitochondria, which are believed derived from "purple bacteria", alpha-proteobacteria most closely related to Rickettsiales.
The cpDNA genes encode some of the molecules needed for chloroplast function. Hundreds of others are transcribed from genes in cellular nucleus, translated into proteins in the cytoplasm, and transported into the chloroplast. It has been experimentally demonstrated that endosymbiotic engulfment of prokaryotes is followed by Endosymbiotic Gene Transfer.
Chloroplast RNA-binding and pentatricopeptide repeat proteins
Chloroplast gene expression is mainly regulated at the post-transcriptional level by numerous nuclear-encoded RNA-binding protein factors. In the present study, we focus on two RNA-binding proteins: cpRNP (chloroplast ribonucleoprotein) and PPR (pentatricopeptide repeat) protein. These are suggested to be major contributors to chloroplast RNA metabolism. Tobacco cpRNPs are composed of five different proteins containing two RNA-recognition motifs and an acidic N-terminal domain. The cpRNPs are abundant proteins and form heterogeneous complexes with most ribosome-free mRNAs and the precursors of tRNAs in the stroma. The complexes could function as platforms for various RNA-processing events in chloroplasts. It has been demonstrated that cpRNPs contribute to RNA stabilization, 3´-end formation and editing. The PPR proteins occur as a superfamily only in the higher plant species. They are predicted to be involved in RNA/DNA metabolism in chloroplasts or mitochondria. Nuclear-encoded HCF152 is a chloroplast-localized protein that usually has 12 PPR motifs. The null mutant of Arabidopsis, hcf152, is impaired in the 5´-end processing and splicing of petB transcripts. HCF152 binds the petB exon–intron junctions with high affinity. The number of PPR motifs controls its affinity and specificity for RNA. It has been suggested that each of the highly variable PPR proteins is a gene-specific regulator of plant organellar RNA metabolism.
T. Nakamura, G. Schuster, M. Sugiura and M. Sugita Chloroplast RNA-binding and pentatricopeptide repeat proteins Biochem. Soc. Trans.. (2004) 32, (571–574)
mitochondria
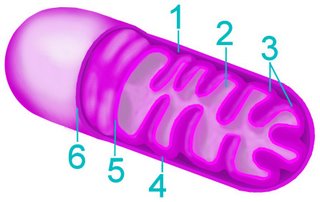 The mitochondrion (pl. mitochondria) is the 'power house of the eukaryotic cell, performing oxidative phosporylation. Mitochondria have two internal, membrane-bound spaces, unlike chloroplasts, which have three internal spaces.
The mitochondrion (pl. mitochondria) is the 'power house of the eukaryotic cell, performing oxidative phosporylation. Mitochondria have two internal, membrane-bound spaces, unlike chloroplasts, which have three internal spaces.The outer mitochodrial membrane is similar in constitution to the eukaryotic cell’s plasma membrane, while the inner membrane is similar in chemical composition to bacterial membranes. This difference is one of several lines of evidence for the serial endosymbiotic origin of mitochondria as phagocytozed purple bacteria.
Above left – click to enlarge : simplified diagram of a mitochondrion showing:
1. space between inner and outer membranes
2. matrix
3. christae
4. junction between membranes
5. inner membrane
6. outer membrane
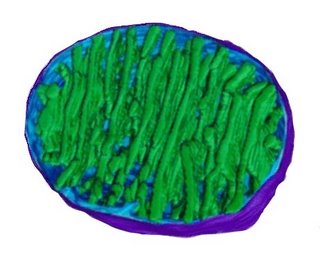
The image at right (click to enlarge) is based on reconstruction of serial tem slices through a mitochondrion. The outer membrane (violet) surrounds the organelle. The inner membrane (pale blue) is contiguous, at membrane junctions (pale blue connecting to green at lower center), with the inner membrane that forms the walls of cristae (green).
The matrix – a soup factory – lies between the cristae, and contains mitochondrial DNA and the components of intermediary metabolism. image - mitochondrion cut : tour mitochondrion :
The outer and inner membranes are composed of phospolipid bilayers studded with proteins, much like the cell membrane. However, the composition of the inner and outer membranes is very different.
The inner mitochondrial membrane contains more than 100 different polypeptides. The protein to phospholipid ratio is very high – more than 3:1 by weight, having about 1 protein for 15 phospholipids. The inner membrane is also rich in an unusual phospholipid, cardiolipin, which is usually characteristic of bacterial plasma membranes. This composition, along with other evidence, has led to the assumption that the inner membrane is derived from endosymbiotic prokaryotes. The endosymbiotic theory of eukaryotic evolution is now widely accepted.
In contrast, the outer membrane, which encloses the entire mitochondrion, is similar in composition to the cell membrane and comprises about 50% phospolipids by weight and contains a variety of enzymes. The enzymes carry out activities such as the oxidation of epinephrine (adrenaline), the degradation of tryptophan, and the elongation of fatty acids.
The plant chloroplast is the site of photosynthesis : animation - chloroplast : tour the chloroplast : Virtual Cell Textbook - Cell Biology







































